Real-Q 2019-nCoV Detection Kit
INSTRUCTIONS FOR USE
(Ver. 1.1, Nov. 2020)
For Use Under Emergency Use Authorization (EUA) Only
For Prescription Use Only
For in vitro Diagnostic Use Only
Intended Use
The Real-Q 2019-nCoV Detection Kit is a real-time RT-PCR test intended for the qualitative detection of nucleic acid from the SARS-CoV-2 in Nasopharyngeal (NP) swab, NP aspirates, nasal mid-turbinate swab, nasal swab, oropharyngeal (throat) swab, sputum, tracheal aspirates, and bronchoalveolar lavage (BAL) specimens from individuals who are suspected of COVID-19 by their healthcare provider. Testing is limited to laboratories certified under the Clinical Laboratory Improvement Amendments of 1988 (CLIA), 42 U.S.C. §263a, which meets the requirements to perform high complexity tests.
Results are for the identification of SARS-CoV-2 RNA. The SARS-CoV-2 RNA is generally detectable in respiratory specimens during the acute phase of infection. Positive results are indicative of the presence of SARS-CoV-2 RNA. Clinical correlation with patient history and other diagnostic information is necessary to determine patient infection status. Positive results do not rule out bacterial infection or co-infection with other viruses. The agent detected may not be the definite cause of the disease. Laboratories within the United States and its territories are required to report all positive results to the appropriate public health authorities.
Negative results do not preclude SARS-CoV-2 infection and should not be used as the sole basis for patient management decisions. Negative results must be combined with clinical observations, patient history, and epidemiological information.
The Real-Q 2019-nCoV Detection Kit is intended for use by qualified clinical laboratory personnel specifically instructed and trained in the techniques of real-time PCR assays and in vitro diagnostic procedures. The Real-Q 2019-nCoV Detection Kit is only for use under the Food and Drug Administration’s Emergency Use Authorization.
Product Description
The Real-Q 2019-nCoV Detection Kit is a real-time reverse transcription-polymerase chain reaction (RRT-PCR) test. The 2019-nCoV primer and probe set(s) is designed to detect RNA from the 2019-nCoV in nasopharyngeal (NP) swab, NP aspirates, nasal mid-turbinate swab, nasal swab, oropharyngeal (throat) swab, sputum, tracheal aspirates, and bronchoalveolar lavage (BAL) specimens from patients who are suspected of COVID-19 by their healthcare provider.
The oligonucleotide primers and probes for detection of SARS-CoV-2 were selected from regions of the virus RNA dependent RNA polymerase (RdRP) gene and Envelope (E) gene. An additional primer/probe set to detect the human RNase P gene (HRP) in clinical specimens is also included in the kit. The target gene for detection and the fluorescent dye of the probe are shown in the table below. The test is designed to detect TaqMan probe fluorescence signals in three different wavelengths in a single tube.
| FAM | HEX/VIC | Cy5 |
| RdRP gene | E gene | Human RNase P |
The RdRP primers & probe (FAM dye is attached) detect specific SARS-CoV-2 sequence and the E gene primers & probe (HEX dye is attached) detect both SARS-CoV and SARS-CoV-2 belonging to the B lineage of Betacoronavirus. The HRP-specific probe (internal control, IC) is labeled with a different fluorophore (Cy5 dye is attached), thus allowing for simultaneous detection of both SARS-CoV-2 and IC amplified products in the same reaction well.
Principle of the Procedure
Nucleic acids are isolated and purified using either QIAamp® MinElute Virus Spin Kit (QIAGEN) or MagNA Pure 96 DNA and Viral NA Small Volume Kit (Roche).
The performance of the Real-Q 2019-nCoV detection Kit is dependent upon the amount and quality of template RNA purified from human specimens. The following commercially available RNA extraction kits and instruments have been qualified and validated for recovery and purity of RNA for use with the kit:
| Extraction Kit (Manufacturer) | |
| Manual extraction | QIAamp MinElute Virus Spin Kit (QIAGEN) |
| Automated extraction system (MagNA pure 96) | DNA and Viral RNA Small Volume kit (Roche) |
The purified nucleic acid is reverse transcribed and amplified in a combined reaction process. 5 μL RNA template is added into 20 μL RT-PCR master mixture which is prepared from 12.5 ul of 2x PCR reaction mixture, 3 ul of nCoV probe & primer mixture, and 1 μl of RT-PCR Enzyme. The real-time RT-PCR reaction is performed on Applied Biosystems 7500 Real-Time PCR Instrument System (software version 2.3), Applied Biosystems 7500 fast Real-Time PCR Instrument System (software version 2.3, Applied Biosystems QuantStudio 5 Real-Time PCR Instrument System (software version 1.4), CFX96 real-time PCR detection system (software version 1.6), and CFX96 DX real-time PCR detection system (software version 1.6). In the RT-PCR reaction, the viral RNA is first converted into cDNA in a reverse transcription reaction. The cDNA is then amplified by the target-specific forward and reverse primers in the PCR reaction.
In the process, the probe anneals to a specific target sequence located between the forward and reverse primers. During the extension phase of the PCR cycle, the 5’ nuclease activity of Taq polymerase degrades the probe, causing the reporter dye to separate from the quencher dye, generating a fluorescent signal. With each cycle, additional reporter dye molecules are cleaved from their respective probes, increasing the fluorescence intensity. Fluorescence intensity is monitored at each PCR cycle by Applied Biosystems 7500/7500 fast/QS5 Real-Time PCR Instrument System, Bio-Rad CFX96/CFX96 DX real-time PCR detection system.
Components
| Component | Description | Volume (100T) |
| PCR MIX | 2X PCR reaction mixture | 1,250 µl |
| PROBE | nCoV probe & primer mixture | 300 µl |
| POSITIVE CONTROL | Positive control | 50 µl |
| ENZYME | RT-PCR enzyme | 100 µl |
| WATER | Water, sterile, DNase/RNase fre | 1,000 µl |
| ROX DYE | ROX reference dye | 10 µl |
Note :
- The 2X PCR reaction mixture and RT-PCR enzyme contain reagents and enzymes for reverse transcription and amplification of SARS-CoV-2 targets (RdRP gene and E gene) and human RNase P (HRP).
- The nCoV probe & primer mixture includes RdRP gene detection primer/probe, E gene detection primer/probe, and HRP detection primer/probe.
- The Positive Control will yield a positive result with the RdRP gene, E gene, and HRP primer/probe sets. The Positive Control contains the RdRP gene, Eugene, and HRP gene at a concentration of 160x LoD. The use of an additional Positive Control that is closer to the LoD is recommended as a further QC measure.
- Water is to be used as a “no template”/negative control to check for contamination during processing and must be included in each run.
- ROX is a passive reference dye used in ABI 7500/7500fast/QS5 real-time PCR instruments.
- All components are to be taken out immediately before use, thawed, and used following centrifugation. Immediately after use, store at -20 ℃.
Materials required but not provided
| RNA extraction | • QIAGEN QIAamp MinElute Virus Spin Kit (Cat. No. 57704) |
| • Roche MagNA pure 96/DNA and Viral RNA Small Volume Kit | |
| Real-time PCR instrument | •Applied Biosystems 7500 real-time PCR instrument (7500 Software v2.3) |
| • Applied Biosystems 7500 fast real-time PCR instrument (7500 Software v2.3) | |
| • Applied Biosystems QuantStudio 5 real-time PCR Instrument (QuantStudio™ Design and Analysis Software v1.4) | |
| • CFX96 real-time PCR detection system (CFX Manager™ Software v1.6) | |
| • CFX96 DX real-time PCR detection system (CFX Manager™ Software v1.6) | |
| Consumables and Equipment | • Disposable powder-free gloves • Pipettes (adjustable) and Nuclease-free pipet tips with aerosol barriers • Nuclease-free, low-binding microcentrifuge tubes (1.5 ml) • 0.2 mL DNase-free PCR tubes or plates (96 well) recommended by the instrument manufacturer • Optical caps or optical adhesive cover recommended by the instrument manufacturer • Desktop centrifuge •(1.5 mL microcentrifuge, 8 strip tubes centrifuge, 96 well plate centrifuge) |
Warnings & Precautions
- For in vitro diagnostic use only
- For prescription use only
- For Emergency Use Authorization only.
- This test has not been FDA cleared or approved; the test has been authorized by FDA under an Emergency Use Authorization (EUA) for use by laboratories certified under the Clinical Laboratory Improvement Amendments (CLIA) of 1988, 42 U.S.C. §263a, that meet requirements to perform high complexity tests.
- This test has been authorized only for the detection of nucleic acid from SARS-CoV-2, not for any other viruses or pathogens.
- This test is only authorized for the duration of the declaration that circumstances exist justifying the authorization of emergency use of in vitro diagnostic tests for detection and/or diagnosis of COVID-19 under Section 564(b)(1) of the Act, 21 U.S.C. § 360bbb-3(b)(1), unless the authorization is terminated or revoked sooner.
- All patient specimens and positive controls should be considered infectious and/or biohazardous and handled accordingly with safe laboratory procedures.
- Use personal protective equipment (PPE) consistent with current guidelines for the handling of potentially infectious samples.
- Do not eat, drink, smoke, apply cosmetics or handle contact lenses in areas where reagents and human specimens are handled.
- Do not use products beyond the expiration date.
- Store the kit at -20 °C.
- Do not mix with other product numbers.
- Always wear laboratory gloves, lab coats, and goggles when handling kit components.
- It is recommended to use an aerosol barrier, RNase, DNase-free tips to prevent contamination.
- All components are to be taken out immediately before use, thawed, and used after centrifugation for several seconds.
- After using the kit components, immediately store at -20 ℃, and limit freezing and thawing to 5 times.
- Reagents should be used immediately after thawing to reduce the time at room temperature.
- When dispensing sample RNA, dispense RNA into the designated well.
- Manipulation of strip caps should be performed only while wearing vinyl gloves or rubber gloves without powder.
- If the tube is not well closed with a striped cap, the contents may evaporate, and incorrect results may occur.
- After adding the sample RNA to the appropriate tube, close the strip tube with a stripped cap, lightly centrifuge, and transfer the tube to a real-time PCR instrument.
- Plate covers should be manipulated only while wearing plastic gloves or rubber gloves without powder.
- If the plate is not covered with a plate cover, the contents may evaporate, and incorrect results may occur.
- After completing the sample RNA loading, cover the plate with a plate cover, lightly centrifuge and install the plate on a real-time PCR instrument.
- Check if the real-time PCR conditions and fluorescence dye selection described in the instruction manual are correctly set before proceeding.
- If the positive control is not amplified, the entire run should be repeated from residual extracted RNA as the result is considered invalid.
- If a positive amplification signal appears in the negative control, the entire run should be repeated from residual extracted RNA as the result is considered invalid.
- Verify the amplification curve for each sample to verify that the Ct analysis is correct.
- PCR is a very sensitive method, therefore, take care to avoid carry-over contamination.
- Avoid microbial and ribonuclease contamination of kit reagents including positive and negative controls.
- Discard the positive control DNA immediately once the kit has been consumed.
- Dispose of unused reagents, waste, and samples according to regulations.
- If reagent gets into your eyes, immediately flush extensively with water and follow the doctor’s instructions.
- If the reagent comes into contact with skin, immediately rinse with water.
- When handling samples that may cause infection, treat them safely according to CLSI Guideline M29-A.
- Maintain real-time PCR instruments according to the manufacturer’s instructions.
Reagent Storage
The components of the Real-Q 2019-nCoV Detection Kit should be stored at -20 °C. Under these conditions, components of the kit are stable until the expiry date stated on the label. Multiple freeze-thaw cycles should be avoided and should not exceed five freeze-thaw cycles as this may reduce the analytical sensitivity.
Note: Do not store the kit at room temperature.
Control Materials
Positive Control (PC):
- The positive control is comprised of non-infectious DNA plasmids containing the RdRP gene, E gene, and RNase P gene fragments used to verify the PCR amplification process and is used in every test. The PC should yield a positive result for each target in the Real-Q 2019-nCoV Detection Kit. The Positive Control is at a concentration of 160x LoD. The use of an additional Positive Control that is closer to the LoD is recommended as a further QC measure.
- Negative Control (NC):
The negative control is DNase/RNase free water that is used to monitor non-specific amplification and contamination during the RT-PCR processes. The NC should yield a negative result for each target in the Real-Q 2019-nCoV detection Kit. - Internal Control
The Real-Q 2019-nCoV Detection Kit uses the Human RNase P (HRP) gene as an endogenous extraction control, so RNA can be extracted and tested directly from the sample without the need for further addition of control material. RNase P is used to monitor extraction, reverse transcription, and real-time PCR amplification processes.
Additional controls should be tested in accordance with state and institutional guidelines and accreditation requirements. Testi ng of an RNA-based external positive control (such as a known SARS-CoV-2 positive specimen) with every batch of patient samples (extraction through RT-PCR) is recommended.
Specimen
- Sample collection
Sample collection device and/or sample preservation buffer is not included as part of the kit. Refer to the CDC Interim Guidelines for Collecting, Handling, and Testing Clinical Specimens for COVID-19. https://www.cdc.gov/coronavirus/2019-ncov/lab/guidelines-clinicalspecimens.html Nasopharyngeal (NP) swab, NP aspirates, nasal mid-turbinate swab, nasal swab, oropharyngeal (throat) swab, sputum, tracheal aspirates, and bronchoalveolar lavage (BAL) specimens are acceptable specimen types. The use of UTM is recommended for the transportation of swab specimens.
- After collection, the specimen should be stored at 2-8°C for up to 48 hours after collection.
- If delivery and processing exceed 48 hours, specimens should be transported in dry ice and once in laboratory frozen at -70°C or colder.
- Specimen Transport
For domestic and international shipments, specimens must be packaged, shipped, and transported according to the current edition of the International Air Transport Association (IATA) Dangerous Goods Regulation. Follow shipping regulations for UN 3373 Biological Substance, Category B when sending potential SARS-CoV-2 specimens.
Procedure
Schematic workflow
The use of a uni-directional workflow with separate preparation areas for RNA extraction, assay setup, and amplification is recommended to minimize the risk of contamination.
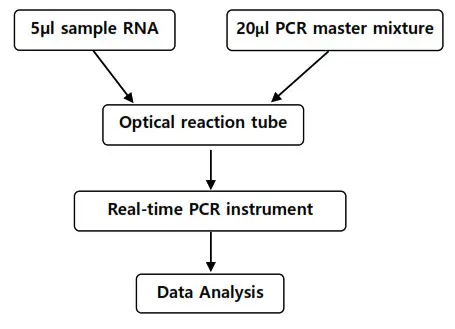
RNA extraction
The following RNA extraction kits have been validated for use with this assay:
| Extraction Kit (Manufacturer) | Sample volume | Elution volume | |
| Manual extraction | QIAamp MinElute Virus Spin Kit (QIAGEN) | 200 µl | 50 µl |
| Automated extraction system (MagNA pure 96) | DNA and Viral RNA Small Volume kit (Roche) | 250 µl | 50 µl |
Viral RNA is extracted according to the manual provided by each manufacturer. The extracted RNA is immediately tested with the Real-Q 2019-nCoV Detection Kit, and if it cannot be tested immediately, it should be stored below -70 ℃.
Preparation of Reaction Mixture
Reagents are to be prepared in the following order:
A. Thaw the components of the kit that are stored at -20 ℃.
B. Keep the components in ice or ice block as soon as they are thawed.
C. Prepare the master mixture in ice or ice block for the PCR.
D. After preparing the master mixture, immediately return kit components to -20 ℃.
E. Mix the master mix gently by pipetting up and down, then centrifuge briefly.
Note: Insufficient mixing of the master mixture may result in incorrect results.
Prepare a real-time PCR master mix
A.Prepare a real-time PCR master mix for the reaction.
- Total required reactions = (n sample +1 positive control +1 negative control) +1
- Preparation of master mixture for CFX96/CFX96 DX
Component Description Volume PCR MIX 2X PCR reaction mixture 12.5 µl PROBE nCoV probe & primer mixture 3 µl ENZYME RT-PCR enzyme 1 µl WATER Water, sterile, DNase/RNase free 3.5 µl 0 Total 20 µl - Preparation of master mixture for AB7500/7500fast/QS5
Component Description Volume PCR MIX 2X PCR reaction mixture 12.5 µl ROX DYE ROX reference dye 0.07 µl PROBE nCoV probe & primer mixture 3 µl ENZYME RT-PCR enzyme 1 µl WATER Water, sterile, DNase/RNase free 3.43 µl Total 20 µl
Note: ROX dye may be added to the 2X PCR reaction mixture before using the kit.
In this case, 12.57 µl of a 2X PCR reaction mixture containing ROX can be used.
Note: When mixing the master mixture, pipette up and down several times; do not vortex.
B. After mixing the master mixture well, briefly centrifuge. Aliquot 20 µl of the master mixture into real-time PCR strip tubes or plates wells.
C. Add 5 µl of the clinical sample RNA into each tube or well-containing aliquot of the master mixture.
D. Add 5 µl of provided positive control (PC) into the PC well-containing aliquot of the master mixture.
E. Add 5 µl of water to the negative control well-containing aliquot of the master mixture.
F. Close the tube caps securely and centrifuge briefly. Cover plate with plate cover and centrifuge briefly.
Real-time PCR Condition
| Step | Temperature | Time | Cycle | Acquisition mode |
| 1 | 50 °C | 30 min | 1 cycle | |
| 2 | 95 °C | 15 min | 1 cycle | |
| 3 | 95ºC | 15 sec | 40 cycles | none |
| 62ºC | 45 sec | Acquiring on FAM, HEX/VIC, Cy5 |
Note :
- Fluorescence is detected at 62°C of step 3 (cycling step).
- In the CFX96/CFX96 DX, the fluorescence is designated by selecting FAM, HEX, and Cy5.
- In the ABI7500/7500fast/QS5, the fluorescence is set FAM, VIC, Cy5 for Reporter, and set all Quencher to None.
| Target | Fluorescence |
| RdRP gene | FAM |
| E gene | HEX/VIC |
| IC(HRP) | Cy5 |
Results analysis
Positive and Negative controls
You should check the Ct value of the positive control. If the Ct value is out of the specified range, re-testing of all samples and controls within the PCR run should be performed. Positive results for the negative control, render the entire run invalid and all samples and controls within the run should be re-tested starting from RT-PCR.
| FAM Ct (RdRP gene) | HEX/VIC Ct (E gene) | Cy5 Ct (HRP) | Result | Comment | |
| Positive control | 28±5 | 28±5 | 28±5 | Positive | Valid |
| Negative control | Neg | Neg | Neg | Negative | Valid |
Interpretation of results
Assessment of clinical specimen test results should be performed after the positive and negative controls have been examined and determined to be valid and acceptable. If the controls are not valid, the patient results cannot be interpreted.
- Analyses should be performed separately for each target using a manual threshold setting.
- Thresholds should be adjusted to fall within the exponential phase of the fluorescence curves and above any background noise signal. The procedure chosen for setting the threshold should be used consistently.
- Threshold setting
Threshold FAM (RdRP gene) HEX / VIC (E gene) Cy5 (HRP gene) CFX96 300 300 200 CFX96 DX 300 300 200 ABI 7500 (with ROX) 0.1 0.05 0.05 ABI 7500 fast (with ROX) 0.1 0.05 0.05 ABI QS5 (with ROX) 0.1 0.05 0.05 - Cut-off Ct is as shown in the table below.
FAM (RdRP gene) HEX / VIC (E gene) Cy5 (HRP gene) ≤38 ≤38 ≤35 To be valid, all clinical samples should exhibit fluorescence growth curves in the RNase P reaction that cross the threshold line within 35 cycles (≤35 Ct).
- Analyze the results according to the table below.
| FAM (RdRP) | HEX/VIC(E gene | Cy5 (HRP) | Result | Comment |
| + | + | +/- | 2019-nCoV positive | |
| + | – | +/- | 2019-nCoV positive (*) | |
| – | + | +/- | Presumptive positive for 2019-nCoV (**) | A retest is recommended. |
| – | – | + | Negative | |
| – | – | – | Invalid | repeat the test |
(*) RdRP gene (+) and / E gene (-) result could be caused by 1) a sample at concentrations near or below the limit of detection of the test, 2) a mutation in the E gene target region in the oligonucleotide binding sites, or 3) other factors. (**) RdRP gene (-) and / E gene (+) result could be caused by 1) a sample at concentrations near or below the limit of detection of the test, 2) a mutation in the RdRP target region in the oligonucleotide binding sites, or 3) infection with some other human coronavirus (e.g., SARS-CoV or another human coronavirus previously unknown to infect humans), or 4) other factors. A retest is recommended. If the repeated result remains presumptive positive, contact your local public health laboratory or CDC for further guidance. Repeated presumptive positive results should be confirmed if clinically needed.
Limitations
- The use of this assay as an in vitro diagnostic under the FDA Emergency Use Authorization (EUA) is limited to laboratories that are certified under the Clinical Laboratory Improvement Amendments of 1988 (CLIA), 42 U.S.C. § 263a, that meet requirements to perform high complexity tests.
- Use only by personnel trained in the techniques of RRT-PCR and in vitro diagnostic procedures.
- Based on the in silico analysis, SARS-CoV and other SARS-like coronaviruses in the same subgenus (Sarbecovirus) as SARS-CoV-2 may cross-react with the E-gene target. SARS-CoV is not known to be currently circulating in the human population, therefore is highly unlikely to be present in patient specimens.
- Specimens must be collected, transported, and stored using appropriate procedures and conditions. Improper collection, transport, or storage of specimens may hinder the ability of the assay to detect the target sequences and lead to false negative results.
- Extraction and amplification of nucleic acid from clinical specimens must be performed according to the specified methods listed in this procedure. Other extraction approaches and processing systems have not been evaluated.
- Avoid contamination by adhering to good laboratory practices and to the procedures specified in this package insert.
- Laboratories are required to report all positive results to the appropriate public health authorities.
- Detection of SARS-CoV-2 RNA may be affected by sample collection methods, patient factors (eg, presence of symptoms), and/or stage of infection.
- Positive and negative predictive values are highly dependent on prevalence. False-negative test results are more likely when the prevalence of the disease is high. False-positive test results are more likely when prevalence is moderate to low
- Negative results do not preclude infection with the SARS-CoV-2 virus and should not be the sole basis of a patient treatment/management or public health decision. Follow-up testing should be performed according to the current CDC or public health agency recommendations.
- Detection of viral RNA may not indicate the presence of an infectious virus or that SARS-CoV-2 is the causative agent for clinical symptoms.
- The performance of this test has not been established for monitoring the treatment of SARS-CoV-2 infection.
- Do not use expired reagents.
The Real-Q 2019-nCoV Detection Kit Letter of Authorization, along with the authorized Fact Sheet for Healthcare Providers, the authorized Fact Sheet for Patients, and authorized labeling are available on the FDA website: https://www.fda.gov/medical-devices/coronavirus-disease-2019-covid-19-emergency-use-authorizations-medical-devices/vitro-diagnostics-euas. However, to assist clinical laboratories using the Real-Q 2019-nCoV Detection Kit, the relevant Conditions of Authorization are listed below:
A. Authorized laboratories 1 using the Real-Q 2019-nCoV Detection Kit will include with test result reports, all authorized Fact Sheets. Under exigent circumstances, other appropriate methods for disseminating these Fact Sheets may be used, which may include mass media.
B. Authorized laboratories using the Real-Q 2019-nCoV Detection Kit will use the Real-Q 2019nCoV Detection Kit as outlined
in the “Real-Q 2019-nCoV Detection Kit Instructions for Use”. Deviations from the authorized procedures, including the authorized instruments, authorized extraction methods, authorized clinical specimen types, authorized control materials, authorized other ancillary reagents, and authorized materials required to perform the Real-Q 2019-nCoV Detection Kit are not permitted. C. Authorized laboratories that receive the Real-Q 2019-nCoV Detection Kit must notify the relevant public health authorities of their intent to run the test prior to initiating testing.
D. Authorized laboratories using the Real-Q 2019-nCoV Detection Kit will have a process in place for reporting test results to healthcare providers and relevant public health authorities, as appropriate.
E. Authorized laboratories will collect information on the performance of the test and report to DMD/OHT7-OIR/OPEQ/CDRH (via email: [email protected]) and BioSewoom Inc. (via telephone: +82-2-498-2340; web address: http://en.biosewoom.com/; email: [email protected]) any suspected occurrence of false positive or false negative results and significant deviations from the established performance characteristics of the test of which they become aware.
F. All laboratory personnel using the test must be appropriately trained in PCR techniques and use appropriate laboratory and personal protective equipment when handling this kit, and use the test in accordance with the authorized labeling.
G. BioSewoom Inc., authorized distributors, and authorized laboratories using the Real-Q 2019nCoV Detection Kit will ensure that any records associated with this EUA are maintained until otherwise notified by FDA. Such records will be made available to
FDA for inspection upon request.
1The letter of authorization refers to, “Laboratories certified under the Clinical Laboratory Improvement Amendments of 1988 (CLIA), 42 U.S.C. §263a, that meet requirements to perform high complexity tests” as “authorized laboratories.”
Performance evaluation
Analytical sensitivity (Limit of Detection, LoD)
To determine the Limit of Detection (LoD) and analytical sensitivity of the kit, studies were performed using serial dilutions of SARS-CoV-2 RNA and the LoD was determined to be the lowest concentration of template that could reliably be detected in 95% of all positive replicates. After preliminary studies using IVT RNA, the LoD of each target assay in the Real-Q 2019-nCoV Detection Kit was determined and verified using SARS-CoV-2 genomic RNA (RNA sample from the National Culture Collection for Pathogen (NCCP) in the Republic of Korea. NCCP No. 43326). Whole viral genome RNA was spiked into sputum at various concentrations (3.125 copies/µl, 6.25 copies/µl, 12.5 copies/µl) diluted from the stock concentration of 6×10 8 copies/µl. 20 replicates were tested from extraction to real-time RT-PCR analysis. RNA extraction was performed by automatic extraction using the Roche MP96 instrument and manual extraction using the QIAGEN QIAamp MinElute Virus Spin Kit. The real-time RT-PCR assay was performed on ABI 7500 instrument. The LoD in sputum was confirmed as 6.25 copies/µl for both extraction methods.
Additional testing to confirm the LoD with nasopharyngeal swabs and sputum for the remaining thermocyclers was performed using QIAGEN QIAamp MinElute Virus Spin Kit. Following tentative LoD testing on all devices, LoD of the Real-Q 2019-nCoV Detection Kit test was confirmed by testing 20 replicates. The final confirmed LoD test results derived using the QIAGEN QIAamp MinElute Virus Spin Kit for extraction are summarized in the table below.
| ABI7500 | ABI7500 fast | QS5 | CFX96 | CFX96 DX | ||
| Final LOD | Sputum (+) rate | 6.25 copies/pi (100%, 20/20) | 6.25 copies/pi (100%, 20/20) | 6.25 copies/pi (100%, 20/20) | 6.25 copies/pi (95%, 19/20) | 6.25 copies/pi (100%, 20/20) |
| NPS (+) rate | 6.25 copies/pi (100%, 20/20) | 3.125 copies/pi (95%, 19/20) | 6.25 copies/pi (100%, 20/20) | 3.125 copies/pi (100%, 20/20) | 6.25 copies/pi (100%, 20/20) |
FDA SARS-CoV-2 Reference Panel Testing
The evaluation of sensitivity and MERS-CoV cross-reactivity was performed using reference material (T1), blinded samples, and a standard protocol provided by the FDA. The study included a range-finding study and a confirmatory study for LoD. Blinded sample testing was used to establish specificity and to confirm the LoD. The extraction method and instrument used were QIAGEN QIAamp MinElute Virus Spin Kit and CFX96 real-time PCR detection system. The results are summarized in Table 1.
Table 1: Summary of LoD Confirmation Result using the FDA SARS-CoV-2 Reference Panel
| Reference Materials Provided by FDA | Specimen Type | Product LoD | Cross-Reactivity |
| SARS-CoV-2 | NPS | 5,400 NDU/mL | N/A |
| MERS-CoV | N/A | ND |
- NDU/mL = RNA NAAT detectable units/mL
- N/A: Not applicable
- ND: Not detected
Inclusivity (Analytical Sensitivity)
Inclusivity was demonstrated by comparing the Real-Q 2019-nCoV Detection Kit primers and probes to an alignment of 2,753 SARS-CoV-2 sequences downloaded from the NCBI database and 21,587 SARS-CoV-2 sequences downloaded from the GISAID database as of June 1, 2020. Multiple sequence alignment by The MUSCLE alignment software was performed for all SARSCoV-2 sequences. The inclusivity of 2019-nCoV detection was confirmed by in silico analysis of the position of the detection probe and primers used in the Real-Q 2019-nCoV Detection Kit in the alignment data through the GeneDoC MSA program. The genetic multiple sequence alignment of primer and probe sequences showed that the RdRP primer/probe set and E primer/probe set in the Real-Q 2019-nCoV Detection Kit had 100% homology to all the SARS-CoV-2 sequences.
Cross-reactivity (Analytical Specificity)
Cross-reactivity of the Real-Q 2019-nCoV Detection Kit was evaluated by in silico analysis and by wet testing whole organisms and purified nucleic acids from pathogens that may potentially be found in respiratory specimens.
For the in silico analysis, the Real-Q 2019-nCoV Detection Kit primer and probe sequences were queried individually against sequences of 32 pathogens potentially present in upper respiratory specimens and/or with genetic similarities to SARS-CoV-2. Results of in silico analysis demonstrate that there is significant homology between the SARS-coronavirus (NC_004718.3) and our assay primer/probes for the RdRP gene and E gene. The primers and probe for the E gene have 100% homology to the SARS-coronavirus sequence. Upon further analysis of the subject sequence, three single nucleotide mismatches were found in the RdRP probe binding region resulting in an alignment of 22/25 bases. In addition, three single nucleotide mismatches were found in the 3′-terminal and mid-position sequences of the RdRP reverse primer binding region, resulting in an alignment of 17/20 bases. Therefore, it is not anticipated that SARS coronavirus will be detected by the RdRP primer/probe set. SARS-CoV was confirmed not to be amplified by RdRP primer, probe in Real-Q 2019-nCoV Detection kit test through wet testing (see below).
| Pathogen | %Homology Test RdRP gene primer&probe | %Homology Test E gene primer&probe |
| Human Coronavirus 229E | None | None |
| Human Coronavirus 0C43 | None | None |
| Human Coronavirus NL63 | None | None |
| Human coronavirus HKU1 | None | None |
| SARS-coronavirus | 89.% | 100% |
| MERS-coronavirus | None | None |
| Human Adenovirus | None | None |
| Human Enterovirus | None | None |
| Human Influenza A virus | None | None |
| Human Influenza B virus | None | None |
| Human Rhinovirus | None | None |
| Human Parainfluenza virus 4 | None | None |
| Human Parainfluenza virus 3 | None | None |
| Human Parainfluenza virus 2 | None | None |
| Human Parainfluenza virus 1 | None | None |
| Human Respiratory syncytial virus A | None | None |
| Human Respiratory syncytial virus B | None | None |
| Human Metapneumovirus | None | None |
| Human Bocavirus | None | None |
| MycopLAsma pneumonia | None | None |
| Chlamydia pneumoniaae | None | None |
| Streptococcus pneumonia | None | None |
| Haemophilus Influenzae | None | None |
| Legionella pneumophlla | None | None |
| Bordetella pertussis | None | None |
| Mycobacterium tuberculosis | None | None |
| Streptococcus pyogenes | None | None |
| Pneumocystisjirovecii | None | None |
| Candida albicans | None | None |
| Pseudomonas aeruginosa | None | None |
| Staphylococcus epidermis | None | None |
| Staphylococcus salaries | None | None |
Thirty organisms and viruses were wet-tested for cross-reactivity with the Real-Q 2019-nCoV Detection kit. The organisms and viruses were tested by spiking into a sputum matrix at a concentration of 1×10 6 copies/µl. RNA was extracted using the QIAamp MinElute Virus Spin Kit and the Real-Q 2019-nCoV Detection Kit test was performed in duplicate on the AB 7500 Fast Real-Time PCR Instrument. Aside from SARS-Coronavirus yielding positive results with the E gene primer/probe set as predicted through in silico testing, all potentially cross-reactive specimens tested negative with the Real-Q 2019-nCoV Detection Kit, therefore, no further crossreactivity was observed among the wet-tested pathogens.
| NO. | Pathogens | Strain Number | Real-Q 2019-nCoV Detection Kit test results | |
| RdRP gene N=2 | E gene N=2 | |||
| 1 | Adenovirus | ATCC VR-1603 | Negative | Negative |
| 2 | Enterovirus | ATCC VR-1775 | Negative | Negative |
| 3 | Influenza A virus | ATCC VR-1894 | Negative | Negative |
| 4 | Influenza B virus | ATCC VR-101 | Negative | Negative |
| 5 | Coronavirus 299E | ATCC VR-740 | Negative | Negative |
| 6 | Coronavirus 0C43 | ATCC VR-1558 | Negative | Negative |
| 7 | Coronavirus NL63 | ZeptoMetrix 0810228CF | Negative | Negative |
| 8 | Coronavirus HKU1 | Korean isolate | Negative | Negative |
| 9 | SARS-Coronavirus | Korean isolate | Negative | Positive |
| 10 | Rhinovirus | ATCC VR-283 | Negative | Negative |
| 11 | Parainfluenza virus 4 | ATCC VR-1377 | Negative | Negative |
| 12 | Parainfluenza virus 3 | ATCC VR-93 | Negative | Negative |
| 13 | Parainfluenza virus 2 | ATCC VR-92 | Negative | Negative |
| 14 | Parainfluenza virus 1 | ATCC VR-94 | Negative | Negative |
| 15 | Respiratory syncytial virus A | ATCC VR-1540 | Negative | Negative |
| 16 | Respiratory syncytial virus B | ATCC VR-1580 | Negative | Negative |
| 17 | Metapneumovirus | NIBSC 08-320 | Negative | Negative |
| 18 | Bocavirus | ATCC VR-767 | Negative | Negative |
| 19 | Mycoplasma pneumonia | ATCC 15531 | Negative | Negative |
| 20 | Chlamydia pneumonia | ATCC 53592 | Negative | Negative |
| 21 | Streptococcus pneumonia | ATCC 33400 | Negative | Negative |
| 22 | Haemoph#us influenza | ATCC 33391 | Negative | Negative |
| 23 | Legionella pneumophlla | KCCM 41783 | Negative | Negative |
| 24 | Bordetella pertussis | ATCC 9797 | Negative | Negative |
| 25 | Klebsiella pneumoniae | KCCM 11418 | Negative | Negative |
| 26 | Haemophilus hemolyticlcus | ATCC 33390 | Negative | Negative |
| 27 | Bordetella parapertussis | ATCC BAA-587D-5 | Negative | Negative |
| 28 | Streptococcus mills | KCTC 5650 | Negative | Negative |
| 29 | Haemophilus parainfluenza | KCTC 15417 | Negative | Negative |
| 30 | Streptococcus pleuropneumonia | KCTC 5764 | Negative | Negative |
SARS-CoV showed ≥80% homology with RdRP gene primers & probe in silico analysis, yet the SARS-CoV-2 RdRp gene was not detected by the Real-Q 2019-nCoV Detection kit in laboratory testing. The amount of primer(s)/ probe(s) included in the nCoV probe & primer mixture of the Real-Q 2019-nCoV Detection Kit are in such excess that interference is unlikely.
Clinical Evaluation
A clinical evaluation study was conducted to evaluate the performance of the Real-Q 2019-nCoV Detection Kit test using sputum specimens. A total of 20 contrived positive specimens at approximately 2X LoD and 10 contrived positive specimens at approximately 20X to 100X LoD were tested. Samples were contrived by spiking known concentrations of SARS-CoV-2 genomic RNA (RNA sample from the National Culture Collection for Pathogen (NCCP) in the Republic of Korea) into individual sputum specimens. In addition to the contrived positive specimens, 30 negative specimens were tested. Each sample was extracted using the QIAamp MinElute Virus Spin Kit and tested on the ABI 7500 Fast Real-Time PCR Instrument. There were 30 total samples tested twice at the 2X to 100X LoD levels with all results valid and included in the analysis. There were 30 total negative samples tested with all results valid and included in the analysis.
| SARS-CoV-2 Concentration | Number Tested | Positive Detected | % Detection |
| 2X LoD | 20 | 20 | 100 |
| 20X to 100X LoD | 10 | 10 | 100 |
| Negative | 30 | 0 | 0 |
| PPA (Positive Percent Agreement) : 100% , 95% CI: 88.65-100 % | |||
| NPA (Negative Percent Agreement) : 100% , 95% CI: 88.65-100 % | |||
Note: Repeat testing of the 30 positive samples gave the same results.
Mean Ct values for Contrived Positive Specimens:
| Concentration (# positive/# Tested) | RdRP Gene (FAM) | E Gene (HEXNIC) | HRP Gene (Cy5) | |
| 2X LoD (20/20) | Mean Ct | 36.36 | 36.64 | 30. |
| SD | 0.49 | 0.41 | 0.27 | |
| 20X LoD (3/3) | Mean Ct | 34. | 33.8 | 31. |
| SD | 0.24 | 0.19 | 0.31 | |
| 50X LoD (3/3) | Mean Ct | 32.0 | 32. | 30.55 |
| SD | 0.10 | 0.16 | 0.03 | |
| 75X LoD (2/2) | Mean Ct | 32. | 31.6 | 30.55 |
| SD | 0.06 | 0.12 | 0.37 | |
| 100X LoD (2/2) | Mean Ct | 31. | 31. | 30.8 |
| SD | 0.03 | 0.03 | 0.23 |
Values shown are for the first set of assay replicates at each target level; similar mean and SD values were obtained with both sets of replicates.
For the clinical evaluation study, left-over archived samples from patients with suspected and symptomatic COVID-19 infection were used. The Viral collection, Preservation and Transport Medium Kit (CAT. No UTNFS-3B, Noble Biosciences, Inc. Republic of Korea) had been used for the collection of specimens within the clinical study. These samples were previously tested for SARS-CoV-2 using an FDA-authorized assay and stored at a temperature of -70℃ in a clinical laboratory in South Korea prior to inclusion in this study. Specimen information is as shown in the table below.
| NPS/OPS in UTM* | Sputum | total | |
| COVID-19 (+) | 26 | 21 | 47 |
| COVID-19 (-) | 17 | 9 | 26 |
| total | 43 | 30 | 73 |
*These specimens were stored in one UTM after simultaneously collecting NPS and OPS from the same patient. (NPS: Nasopharyngeal swab, OPS: oropharyngeal (throat) swab, UTM: universal transport medium) The purpose of this clinical study was to evaluate the clinical performance of the Real-Q 2019nCoV Detection Kit. In this study, sample RNA was extracted using Qiagen QIAamp MinElute Virus Spin Kit and real-time RT-PCR was performed using CFX96 real-time PCR detection system. All samples were tested by the Real-Q 2019-nCoV Detection Kit and compared with the results of testing with an FDA-authorized assay. The PPA (Positive Percent Agreement) and NPA (Negative Percent Agreement) results were confirmed as follows.
Results of Testing Sputum Specimens
| Comparator test | ||||
| SARS-CoV-2 Positive | SARS-CoV-2 Negative | Total | ||
| Real-Q 2019- nCoV Detection Kit | 2019-Nov Positive | 21 | 0 | 21 |
| 2019-Nov Negative | 0 | 9 | 9 | |
| Total | 21 | 9 | 30 | |
| PPA (Positive Percent Agreement) : 100% (95% CI, 84.5— 100%) NPA (Negative Percent Agreement) : 100% (95% CI, 70.1-100%) | ||||
Results of Testing NP/OP Swab Specimens
| Comparator test | ||||
| SARS-CoV-2 Positive | SARS-CoV-2 Negative | Total | ||
| Real-Q 2019- nCoV Detection Kit | 2019-Nov Positive | 26 | 0 | 26 |
| 2019-Nov Negative | 0 | 17 | 17 | |
| Total | 26 | 17 | 43 | |
| PPA (Positive Percent Agreement) : 100% (95% CI, 87.1— 100%) NPA (Negative Percent Agreement) : 100% (95% CI, 81.6100%) | ||||
References
1) Zhang Y-Z. Novel 2019 coronavirus genome. Virological. [Accessed 21 Jan 2020]. Available from: http://virological.org/t/novel-2019-coronavirus-genome/319
2) Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR Euro Surveill. 2020 Jan;25(3).
3) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan Emerg Microbes Infect. 2020 Dec;9(1):221- 236
Explanation of symbols
| Symbol | Explanation |
| IVD | In vitro diagnostic medical device |
| LOT | Batch code |
| REF | Catalogue number |
 | Use by |
 | Temperature limitation |
 | Contains sufficient for <n> tests |
| Caution | |
| Consult instructions for use | |
 | Manufacturer |
 | Authorized Representative in the European Community |
 | CE mark |
 | Prescription Use only |
| EAU | Emergency Use Authorization |
Contact Information
 BioSewoom Inc.
BioSewoom Inc.
2F Wooyoung Technocenter, 144 Achasan-ro, Seongdong-gu, Seoul, 04783,
Republic of Korea
TEL: +82-2-498-2340
FAX: +82-2-498-1189
Email: [email protected]