Version 2.0 – November 2021
Version 2.0 – November 2021
Web Service Application Programming Interface Software
User Guide
Web Service Application Programming Interface Software

INTRODUCTION
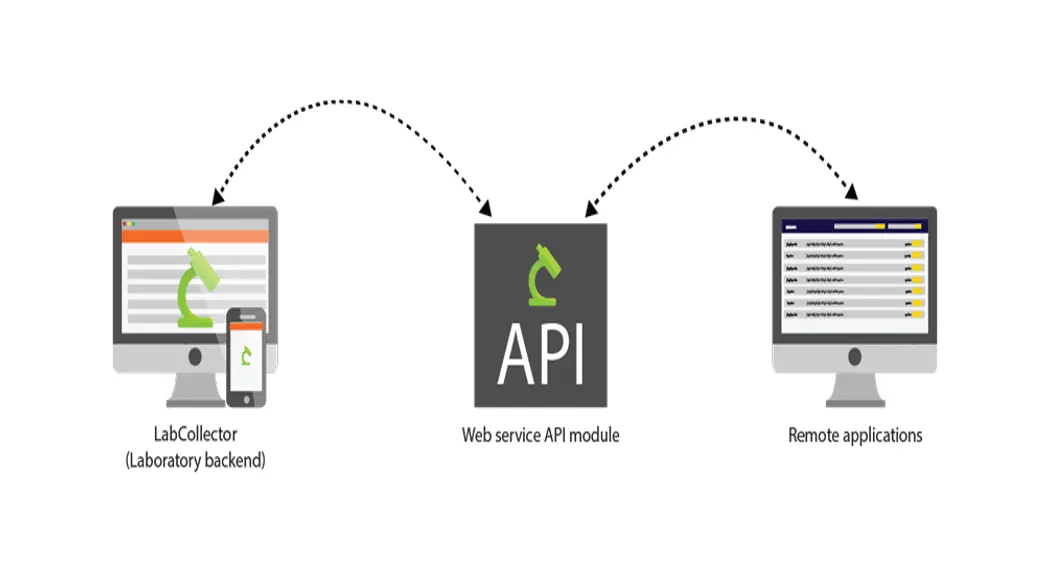
The LabCollector Web Service Application Programming Interface (API) allows third-party applications to interact with LabCollector’s database (modules) and add- ons (ELN and LSM).
The API is based on a Representational State Transfer (REST) architecture allowing access to resources through Uniform Resource Identifier (URI) and actions on them.
Note: Since June 2017 API v1 was discontinued and all new evolutions are in API v2.
LABCOLLECTOR API
2-1. API setup
First of all, you have to declare your application in your LabCollector software. To access the application declaration setup form, log in to LabCollector with super-administrator rights and go to Admin > Setup page. Then select the Web Service API link. 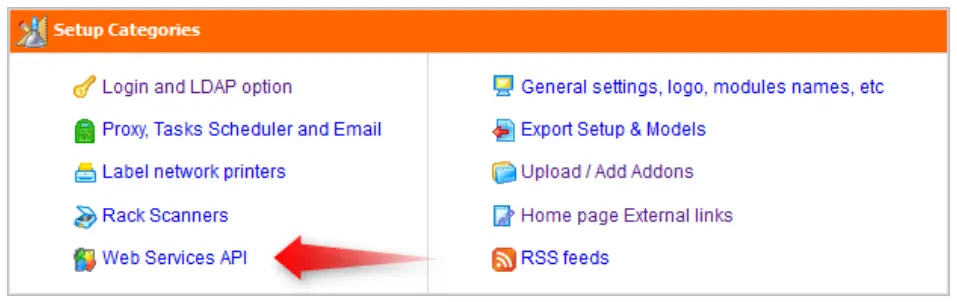 You are now on the Web Service API applications management page. To declare a new application, simply complete this form:
You are now on the Web Service API applications management page. To declare a new application, simply complete this form: 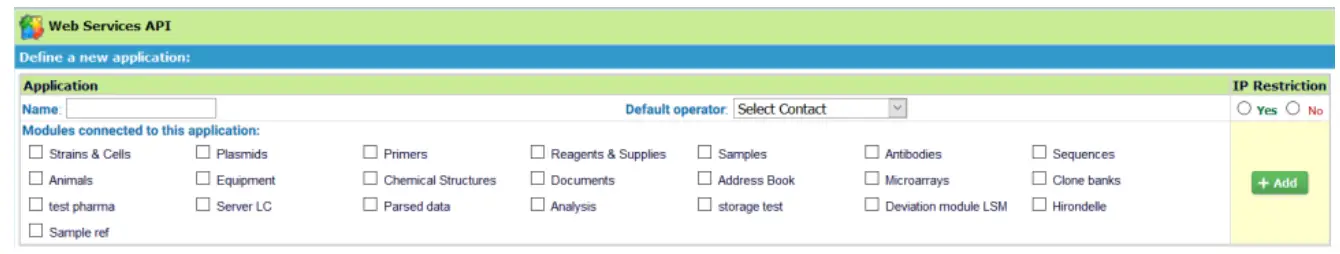
- Name: name of your application.
- Modules connected to this application: select modules in which the application can access.
- Default operator: select the contact that will be the default operator if you don’t want to insert this information in each request.
- IP Restriction: security option allows you to declare a list of IP addresses, which will be allowed you to perform requests on the API.
The Application list shows all the applications for your LabCollector and you can, at any time, modify their scope.
You have also access to the Token which is necessary to identify your application during requests to the API. 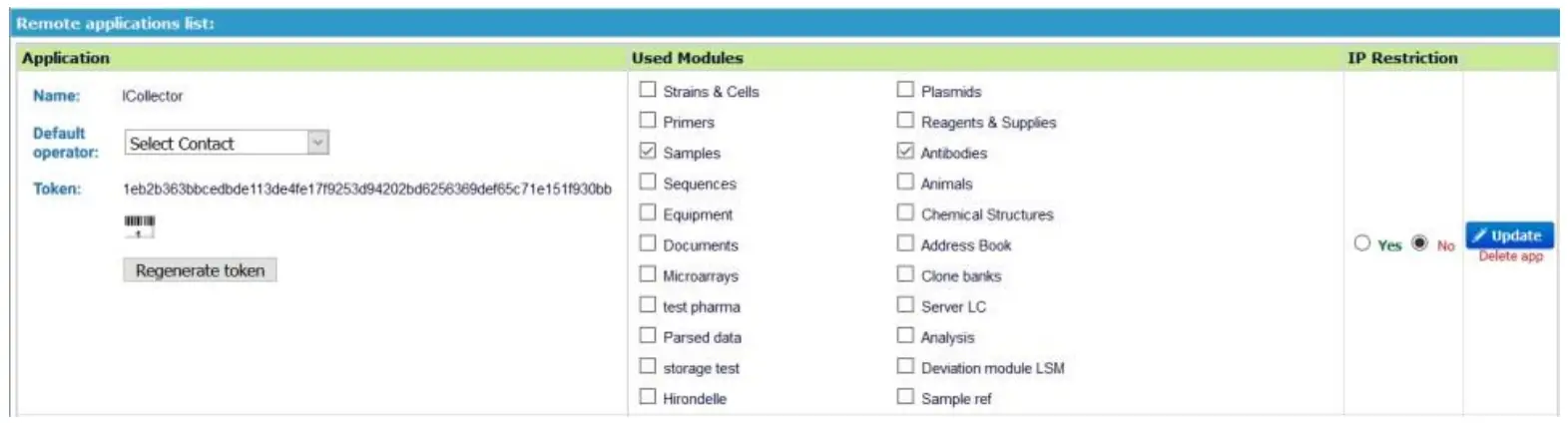
Note: To use this feature, you need to activate Curl on your PHP preferences. In Linux, install the PHP-Curl package.
On windows and with our automatic installer, edit PHP.ini and uncomment extensions for Curl (extension=php_curl.dll).
2-2. Requests
The dialog between third-party applications and the LabCollector web service API is based on the HTTP 1.1 protocol.
2-2-1. API Method
You can send HTTP or HTTPS requests to the web service with a method to act on a resource.
- GET method for reading a resource
- POST method to create a new resource
- PUT method to modify a resource
- DELETE method to delete a resource
2-2-2. Headers
A request to the API requires some specific HTTP/HTTPS headers:
- The Accept header defines the desired response format of your request, text/XML, or application/JSON.
- The X-LC-APP-Auth header is where you put your application token which is necessary to authorize your request to the API.
- The X-LC-APP-Charset header defines the character encoding of your application. It allows the API to send back the response with the appropriate encoding and to correctly convert your POST and PUT requests to the LabCollector’s character encoding (ISO 8859-1).
2-2-3. Tool
You can try to retrieve data from or send data to the API with some software app as Postman (https://www.getpostman.com/).
Uniform Resource Identifier (URI)
2-3-1. GET method
General
Each LabCollector module data is identified by a unique URI (see annex for a complete list of the module’s URI):
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE] This request replies to the list of all data in a module.
You can do searching into module data by adding parameters to your URI. You can pass a parameter with a keyword matching a field value, like:[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?name=[KEYWORD]e.g.
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?name=First%20Record
This request returns the records where their name value contains the “First Record” keyword.
They are some custom parameters that the API uses to perform searching and filtering actions.
Custom parameters
- The record_id parameter to specify data by its ID:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?record_id=[RECORD_ID]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?record_id=1,19
This request returns records with ID 1 and ID 19. You can specify multiple IDs by separating them with a comma.
- The by_keywords parameter performs a keyword search:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?by_keywords=[KEYWORD]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?by_keywords=cell
This request performs a search into all fields of all records and returns matching cells. You can specify multiple keywords by separating them with a comma.
- The by_keywords parameter performs a keyword search:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?by_keywords=[KEYWORD]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?by_keywords=cell
This request performs a search into all fields of all records and returns a matching cell. You can specify multiple keywords by separating them with a comma.
- The fields parameters, If you want to retrieve only some fields values in the API response:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?fields=[FIELD1],[FIELD2]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?fields=count,name
This request returns all records from the module but with only count and name fields. You can specify multiple fields by separating them with a comma.
The request accepts now multiple values separated by a comma, for custom fields of type “select”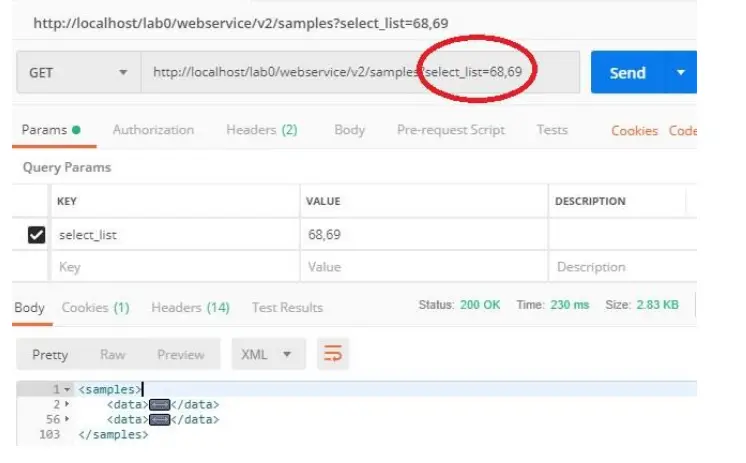
- The search_on parameter allows you to search for data. And you can use it to search by date range as follow:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]&
search_on=date_field&from=XXXXXX&to=ZZZZZZ
If you only use FROM, the result will give all dates bigger than the FROM date. If you only use too, it will return all the value until this date. 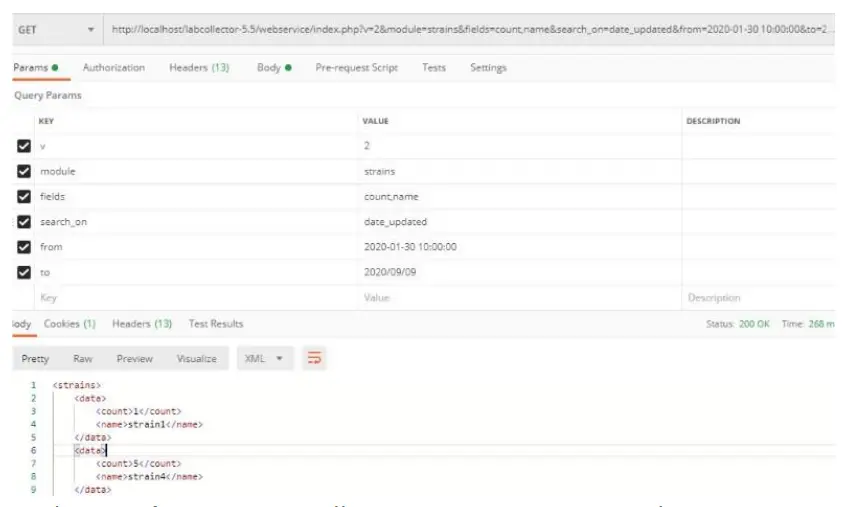
- The sort_by parameter allows you to sort your search:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?sort_by=[FIELD1]_DESC
e.g. [PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?sort_by=name_DESC
This request returns all records sorted in descending order on the name field. You can specify multiple sort_by separating them with a comma and specified order ascendant _ASC” or descendant “_DESC” for each field.
- The limit_to parameter allows you to limit the number of results:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]?limit_to=0,10
This request returns 10 records beginning at index 0. If you do not specify the index, only the number of results indicated is returned.
The API also returns two custom fields in the header response, “X-LC-QUERY-RESULT” containing the number of results returned in the body response and “X-LC-QUERY- TOTAL” containing the total of records matching your search.
Each record also has a unique URI:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]/[DATA_ID]
This request replies a unique record. [DATA_ID] must match the unique ID of the record you want to retrieve.
Storage
You also have Tube Sorter filtering functions for every item linked to storage:
[PATH_TO_LABCOLLECTOR]/webservice/index.php?v=2&action=tube_sorter&box_i d=[BOX_ID]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.php?v=2&action=tube_sorter&box_i d=34
This request returns storage info on box ID 34 like tube sorter. You can specify multiple IDs by separating them with a comma. 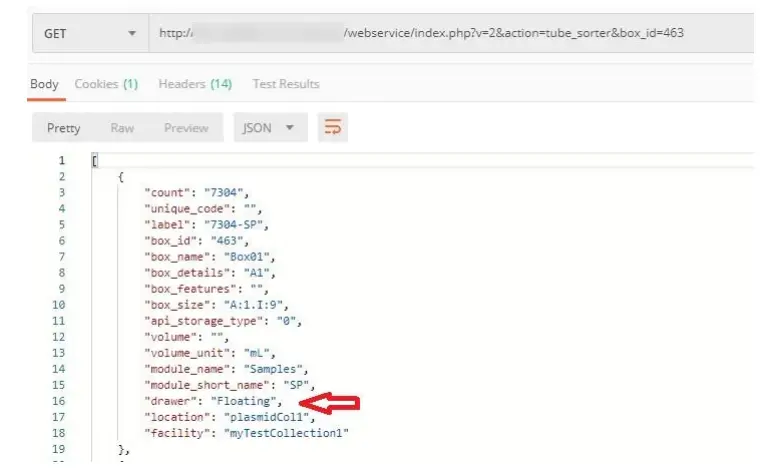
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&box_i d=[BOX_ID]&record_name=[RECORD_NAME]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&box_i d=206&record_name=ST-260
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&recor d_name=[RECORD_NAME]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&recor d_name=ST-260
These requests perform filtering on a record named ST-260. You can specify multiple record names by separating them with a comma. You can also specify box ID, here 206.[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&box_n ame=[BOX_NAME]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=tube_sorter&box_n ame=test-rack_06
This request performs filtering on box test-rack_06. You can specify multiple box names by separating them with a comma.
Other search parameters to action=tube_sorter can be:
- location_id
- location_name
- facility_id
- facility_name
It will return empty boxes as well. - The storage_sec parameter allows retrieving information about secondary storage.
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]&data_id=[DATA_ID]& fields=storage_sec 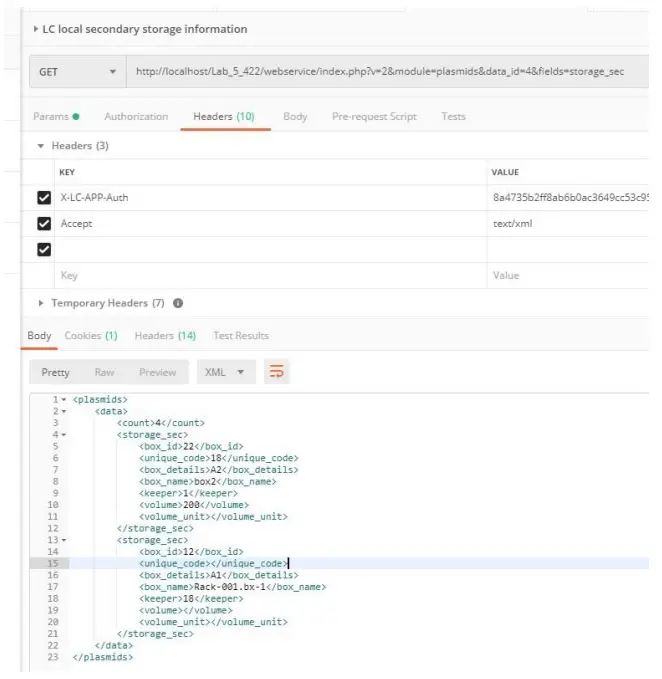
Product lot
- The action get lot allows retrieving lot and reagent info
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getLot
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getLot&lo t_id=1/LT
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getLot&ch em_id=2
Optional parameters are lot_id (in format 1 or 1/LT) and chem_id. If it doesn’t receive parameters, then it retrieves all active lots.
Recipe
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe s
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe &recipe_id=[record_id]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe &recipe_id=509
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe Logs
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe Report&log_id=[record_id]
e.g. [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action=getRecipe Report&log_id=1218
IDs are examples but are mandatory in these calls.
get recipes prints the following info: id, name, description, category 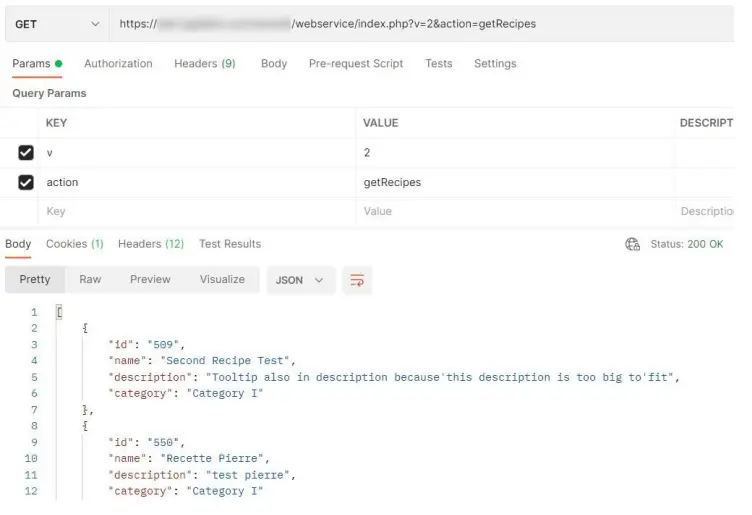
get recipes prints the following info for that recipe_id: id, name, description, category, and then the components 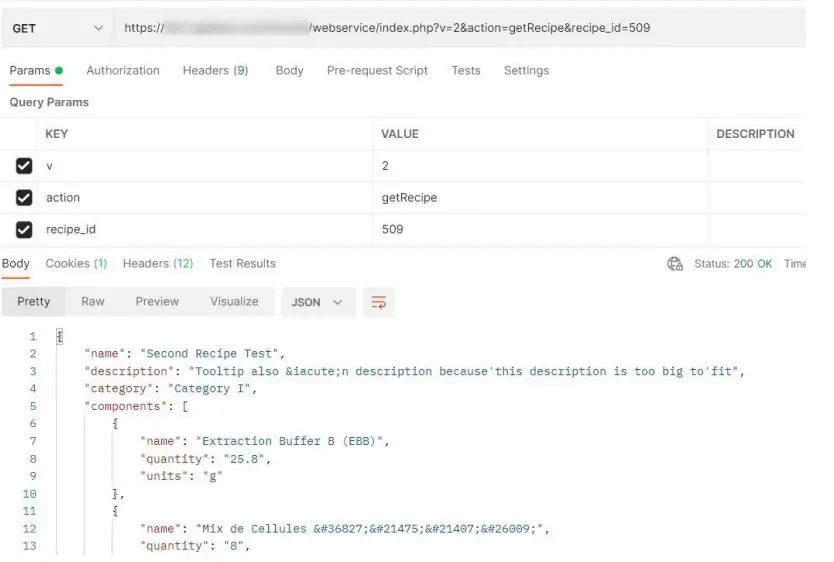 getRecipeLogs prints the following info: id, name, description, category
getRecipeLogs prints the following info: id, name, description, category 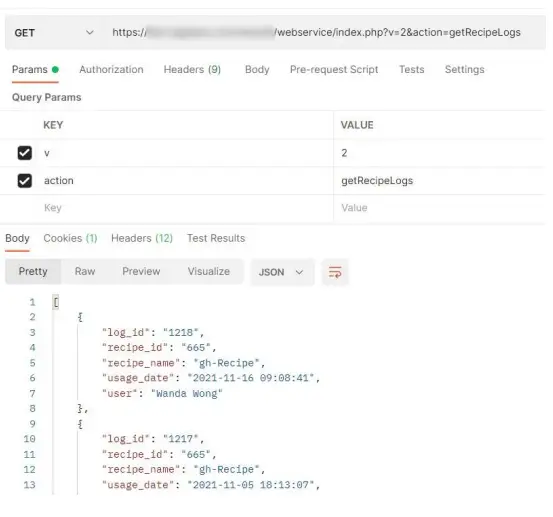 getRecipeReport prints the report PDF for that log_id under the format base64 that can be decoded into PDF.
getRecipeReport prints the report PDF for that log_id under the format base64 that can be decoded into PDF. 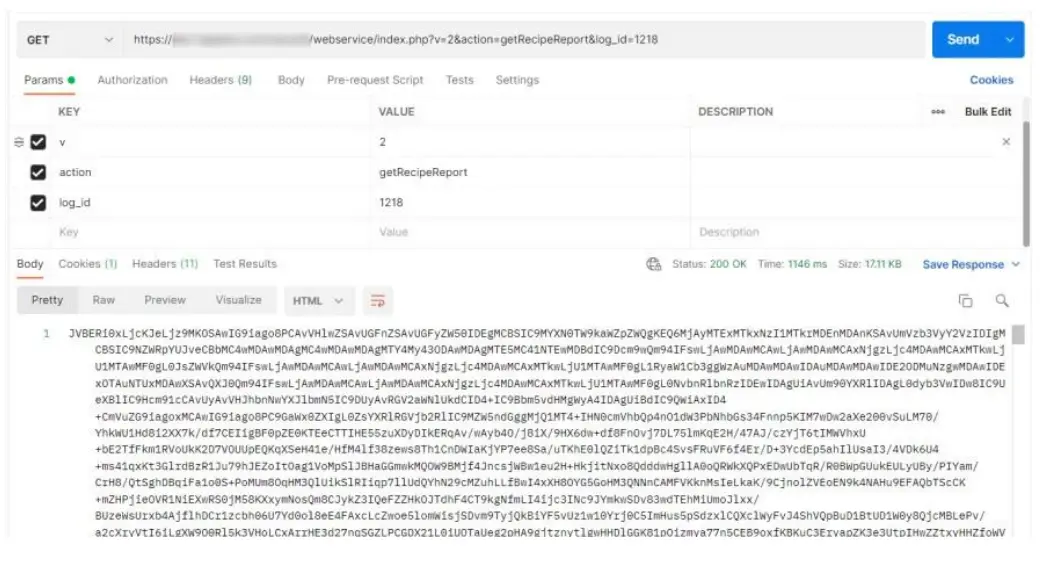
2-3-2. POST method
To create a new resource, simply send a request with the POST method to the desired module URI:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]
Your parameter keys have to match the field’s name.
Check for uniqueness fields that have been added, when creating new records (POST) or update (PUT)
If exists a different record with the same value for a field Uniqueness, API will not complete the action and will return code 409 (Conflict), and the text: Value for field ‘XXX’ must be unique. Value ‘YYY’ already exists in table ‘ZZZ’. (see screenshot) 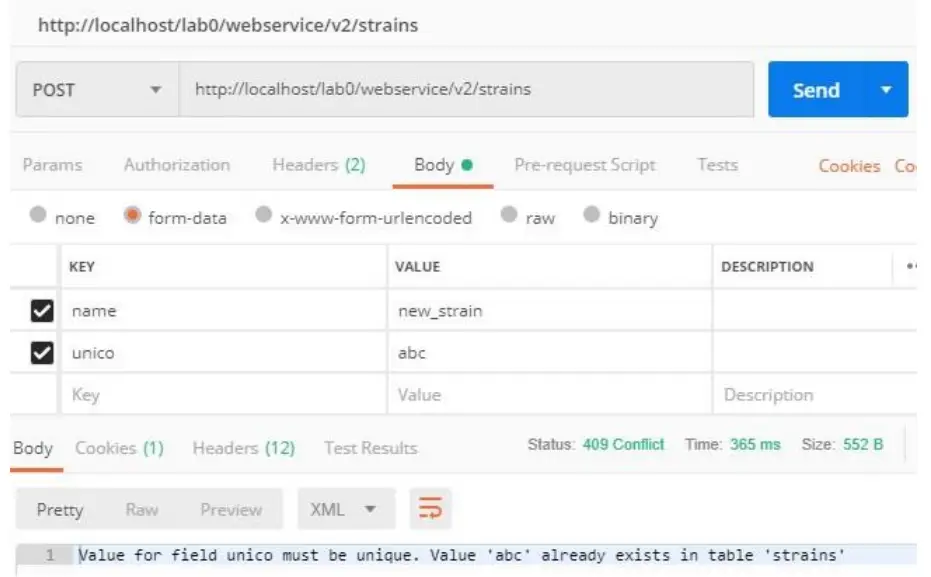
Note: project_code field can be used in POST and PUT and it expects text (not id). You can now create a new project code if it does not exist and if the operator has permissions enough (administrator or super-administrator).
- The action addBox allows you to create a box
[PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2&action= addBox
- Required parameters:
o name
o type (must be a valid type: box, box_nogrid, plate, microplate, visit be, bag, shelf part)
o equipment (supports id or name and must exist in LabCollector storage).
o size (depends on the type of box: should be numeric for a visit be, and the format (A:1.H:8) for a box, a plate, and a microplate) - Optional parameters:
o description
o rack
o keeper
2-3-3. PUT method
To modify a resource, simply send a request with the PUT method to the desired record URI:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]/[DATA_ID]
Your parameter keys have to match the field’s name you want to modify.
For the following actions, note that for PUT requests, parameters must be on the body (not in the URL).
The URL is [PATH_TO_LABCOLLECTOR]/webservice/index.PHP?v=2
The headers are: X-LC-APP-Auth, Accept.
- Remove Volume
– Parameters:
o removeVolume (mandatory)
o barcode, unique_code, or aliquot_barcode (one of them must be present)
o quantity (mandatory)
– Response: OK - Remove Storage
– Parameters:
o remote storage (mandatory)
o barcode, unique_code, or aliquot_barcode (one of them must be present)
– Response: OK - Add Registry Book
– URL:
[PATH_TO_LABCOLLECTOR]/webservice/index.php?v=2&module=[m odule]
– Parameters:
o addRegistryBook (mandatory)
o record_id (mandatory)
o date (mandatory, format yy yy/mm/dd or yyyy-mm-dd)
o comments (mandatory)
o operator (optional, if it does not send the API default operator will be used)
o action (optional, must be a valid ‘Storage Action Type’ defined in LC
>Admin >Preferences > Process & Actions Type)
– Response: OK - Add Secondary storage
– Parameters:
o add secondary storage (mandatory)
o barcode (mandatory)
o box_id (mandatory)
o box_details (mandatory only for the box with grid divider, tube tray, and microplate. If the box is without a grid, a bag, a visit be or a shelf part, it is not required)
o unique_code (optional)
o volume (optional)
o comments (optional)
o cap_color (optional)
Note: An error message is returned if mandatory parameters are not present; if the barcode does not exist; if the unique_code is present, but it isn’t unique; and, if the color is present but it doesn’t exist.
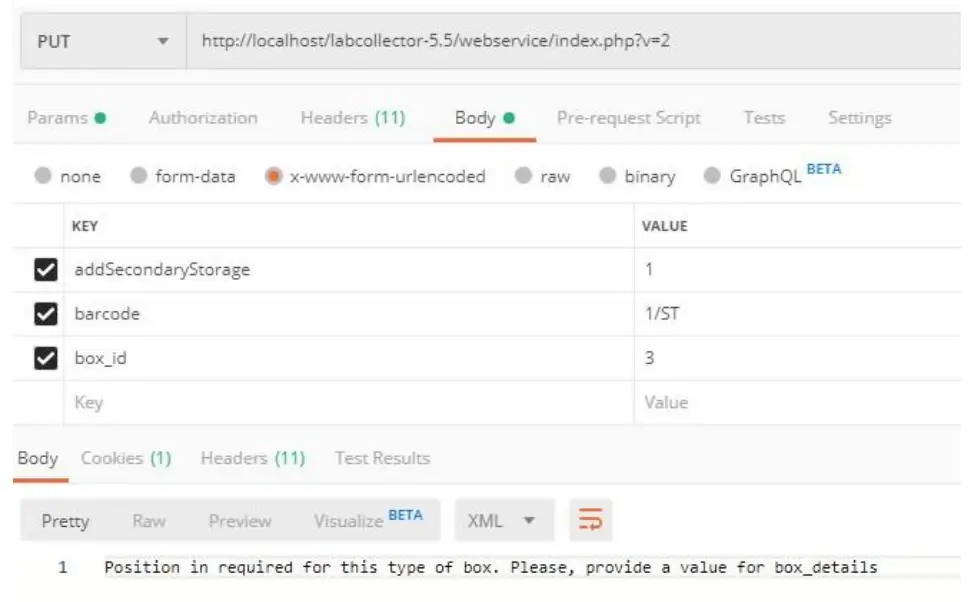
If the parameter box_details is not received and the type of box needs position (box with grid, tube tray, or microplate), an error message is returned. 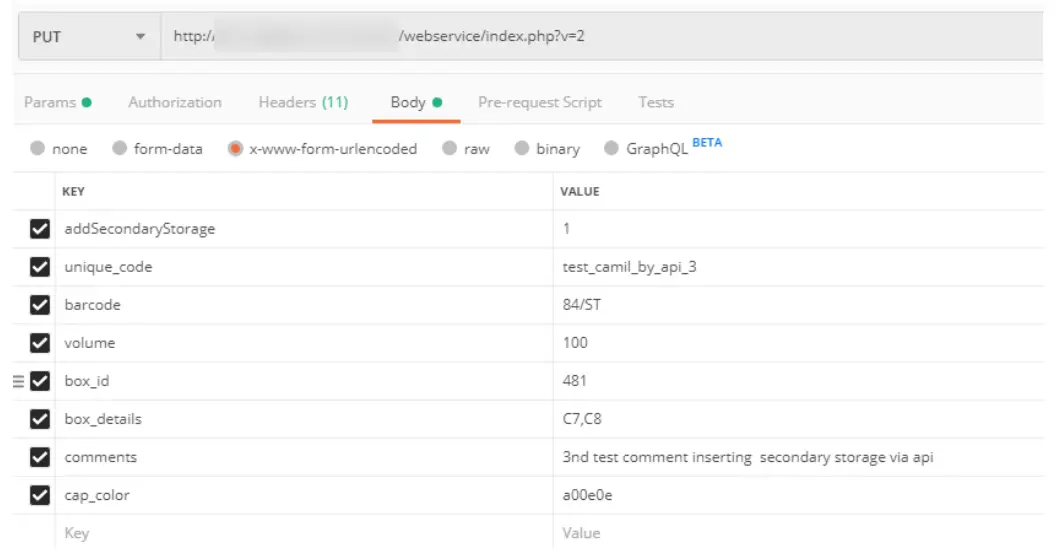
 Note: project_code field can be used in POST and PUT and it expects text (not id). You can now create a new project code if it does not exist and if the operator has permissions enough (administrator or super-administrator).
Note: project_code field can be used in POST and PUT and it expects text (not id). You can now create a new project code if it does not exist and if the operator has permissions enough (administrator or super-administrator).
2-3-4. DELETE method
To delete a resource, simply send a request with the DELETE method to the desired record URI:
[PATH_TO_LABCOLLECTOR]/webservice/v2/[MODULE]/[DATA_ID]
API ERROR MESSAGES
| Message | Response code | Description |
| Requires application authentication to access the Web Service’ | 401 Unauthorized | The request either does not have the header parameter X- LC-APP-Auth or does not have a valid value |
| ‘Invalid Action xxx’ | 400 Bad Request | Parameter action has a different value to ‘tube_sorter’ or ‘NetBackup’ |
| Missing Search Parameters! | 400 Bad Request | The request contains the parameter Action=tube_sorter but it is missing at least one of the following parameters: box_id, box_name, record_name, unique_code, barcode, aliquot_barcode |
| Module “XXX” does not exist!’ | 400 Bad Request | The value of the parameter ‘module’ is not a GB collector module |
| Module “XXX” does not share this data!’ | 403 Forbidden | The value of the parameter ‘module’ is not checked on LabCollector > Admin > Setup > Web service |
| ‘The format of the request is not accepted!’ | 415 Unsupported Media Type | The parameter Accept is used, but the value is not one of the accepted values: application/XML or application/JSON |
| (Empty) | 406 Not Acceptable | The method must be one of the following: GET, POST, PUT, DELETE |
| ‘No data found.’ | 404 Not Found | No data was found with this request’s parameters |
| ‘OK.’ | 200 OK | Record updated successfully |
| ‘Conflict.’ | 409 Conflict | The record could not be updated because there is a conflict in data |
| No organisms value for this module | 404 Not Found | Only the modules “strains”, “samples” and microarrays” have an organism value – you have chosen an incorrect module |
| No categories value for this module | 404 Not Found | Only the module ‘docs’ has categories – you have chosen an incorrect module |
| Webservice requires user authentication | 401 Unauthorized | Deprecated |
| Your IP is not allowed to access this Web Service’ | 401 Unauthorized | The client IP is not in the list of authorized IPs for this Webservices (LC > Admin > Setup > Web service) |
| Error during your request, the following information is mandatory to create a new record: X, Y, Z ‘ | 400 Bad Request | Attempt to post new data without mandatory fields X, Y, Z |
| An error has occurred during your request, the following information is mandatory to remove volume: unique_code or barcode or aliquot_barcode, quantity, quantity | 400 Bad Request | Attempt to remove volume without mandatory parameters: unique_code or barcode or aliquot_barcode, quantity |
| An error has occurred during your request, the following information is mandatory to remove storage: unique_codeor barcode or aliquot_barcode, quantity ‘ | 400 Bad Request | Attempt to remove storage without a mandatory parameter: unique_code or barcode or aliquot_barcode |
| ” | 200 OK | The data requested was returned successfully |
LABCOLLECTOR WEB SERVICE API – ANNEX
The URI system of the API uses a simple and clean URL. Be sure to enable the rewrite engine from Apache to use the URI referenced in the following table. If the LabCollector server does not support the rewrite engine please use the full URL pattern for your request (secondary URL of each line).
| UM | Module | Description | |
| webservice/v2/strains webservice/index.PHP?v=2&module=strai ns | GET POST | Strains & Cells | List of all records |
| webservice/v2/strains/(DATA JD] webservice/index.PHP?v=2&module=strai ns&data jd.[DATA _ID] | GET PUT | Strains & Cells | Unique record |
| webservice/v2/strains/custom fields webservice/index.php?v=2&module=strai ns&getModuleCustomFields=1 | GET | Strains & Cells | List of custom fields |
| webservice/v2/strains/organisms webservice/index.PHP?v=2&module=strai ns&getModuleOrganisms=1 | GET | Strains & Cells organisms | List of |
| webservice/v2/plasmids webservice/index.php?v=2&module=plas mids | GET POST | Plasmids | List of all records |
| webservice/v2/plasmids/IDATAjD] webservice/index.php?v=2&module=plasmids&data _id=IDATA _ID] | GET PUT | Plasmids | Unique record |
| webservice/v2/plasmids/custom fields webservice/index.PHP?v=2&module=plas mids&getModuleCustomFields=1 | GET | Plasmids fields | List of custom |
| webservice/v2/primers webservice/index.PHP?v=2&module=pri mers | GET POST | Primers | List of all records |
| webservice/v2/primers/[DATA JD] webservice/index.PHP?v=2&module=pri mers&data _idADATA _ID] | PUT GET | Primers | Unique record |
| webservice/v2/primers/custom fields | GET | Primers | List of custom fields |
| webservice/index.PHP?v=2&module=pri mers&getModuleCustomFields=1 | |||
| webservice/v2/chemicals webservice/index.PHP?v=2&module=che micals | GET POST | Reagents & Supplies | List of all records |
| webservice/v2/chemicals/IDATA _ID] webservice/index.PHP?v=2&module=che micals&data_idADATA _ID] | GET PUT | Reagents & Supplies | Unique record |
| webservice/v2/chemicals/custom fields webservice/index.PHP?v=2&module=che micals&getModuleCustomFields=1 | GET | Reagents & Supplies fields | List of custom |
| webservice/v2/samples webservice/index.PHP?v=2&module=sam pies | GET POST | Samples | List of all records |
| webservice/v2/samples/IDATA_ID) web service/index.PHP?v=2&module=sam ples&data_id=[DATA _ID] | GET PUT | Samples | Unique record |
| webservice/v2/samples/custom fields webservice/index.PHP?v=2&module=sam ples&getModuleCustomFields=1 | GET | Samples | List of custom fields |
| webservice/v2/samples/organisms webservice/index.php?v=2&module=sam ples&getModuleOrganisms=1 | GET | Samples | List of organisms |
| webservice/v2/samples/types webservice/index.PHP?v=2&module=sam ples&getModuleTypes=1 | GET | Samples | List of sample types |
| webservice/v2/antibodies webservice/index.PHP?v=2&module=anti bodies | GET POST | Antibodies | List of all records |
| webservice/v2/antibodies/(DATA _iDi webservice/index.PHP?v=2&module=anti bodies&data_id=IDATA _ID] | GET PUT | Antibodies | Unique record |
| webservice/v2/antibodies/custom fields webservice/index.PHP?v=2&module=anti bodies&getModuleCustomFields=1 | GET | Antibodies fields | List of custom |
| webservice/v2/sequences webservice/index.PHP?v=2&module=seq uences | GET POST | Sequences | List of all records |
| webservice/v2/sequences/(DATA _iDI webservice/index.PHP?v=2&module=seq uences&data _icHCIATA JD] | GET PUT | Sequences | Unique record |
| webservice/v2/sequences/custom fields webservice/index.PHP?v=2&module=seq uences&getModuleCustomFields=1 | GET | Sequences fields | List of custom |
| webservice/v2/animals webservice/index.PHP?v=2&module=ani mats | GET POST | Animals | List of all records |
| webservice/v2/animals/(DATA JD] webservice/index.PHP?v=2&module=ani mals&data _ick[DATA JD] | GET PUT | Animals | Unique record |
| webservice/v2/animals/custom fields webservice/index.PHP?v=2&module=ani malsketModuleCustomFields=1 | GET | Animals | List of custom fields |
| webservice/v2/equipments webservice/index.php?v=2&module=equi pments | GET POST | Equipment | List of all records |
| webservice/v2/equipments/PATA _el Webservice/index.php?v=2&module=equi pments&data _idADATA _ID] | GET PUT | Equipment | Unique record |
| webservice/v2/equipments/custom fields webservice/index.PHP?v=2&module=equi pments&getModuleCustomFields=1 | GET | Equipment fields | List of custom |
| webservice/v2/structures webservice/index.PHP?v=2&module=stru cures | GET POST | Chemical Structures | List of all records |
| webservice/v2/structures/(DATA_ID] webservice/index.PHP?v=2&module=stru ctures&data jd=(DATA JD] | GET PUT | Chemical Structures | Unique record |
| webservice/v2/structures/custom fields webservice/index.PHP?v=2&module=stru cturesketModuleCustomFields=1 | GET | Chemical Structures | List of custom fields |
| webservice/v2/docs webservice/index.PHP?v=2&module=docs | GET POST | Documents | List of all records |
| webservice/v2/docs/(DATA JD] webservice/index.PHP?v=2&module=docs &data _idADATA _ID] | GET PUT | Documents | Unique record |
| webservice/v2/docs/custom fields webservice/index.php?v=2&module=docs &getModuleCustomFields=1 | GET | Documents | List of custom fields |
| webservice/v2/docs/categories webservice/index.PHP?v=2&module=docs &getModuleCategories=1 | GET | Documents categories | List of |
| webservice/v2/book webservice/index.PHP?v=2&module=abo ok | GET POST | Address Book | List of all records |
| webservice/v2/book/(DATA _ID] webservice/index.php?v=2&module=abo ok&data_idADATA _ID] | GET PUT | Address Book | Unique record |
| webservice/v2/book/custom fields webservice/index.PHP?v=2&module=abo ok&getModuleCustomFields=1 | GET | Address Book | List of custom fields |
| webservice/v2/book/categories webservice/index.PHP?v=2&module=abo ok&getModuleCategories=1 | GET | Address Book categories | List of |
| webservice/v2/microarrays webservice/index.PHP?v=2&module=micr arrays | GET POST | Microarrays | List of all records |
| webservice/v2/microarrays/(DATA_ID] webservice/index.PHP?v=2&module=micr oarrays&data_id=[DATA _ID] | GET PUT | Microarrays | Unique record |
| webservice/v2/microarrays/custom fields webservice/index.PHP?v=2&module=micr oarrays&getModuleCustomFields=1 | GET | Microarrays | List of custom fields |
| webservice/v2/microarrays/organisms webservice/index.PHP?v=2&module=micr oarrays&getModuleOrganisms=1 | GET | Microarrays organisms | List of |
| webservice/v2/(CUSTOM_MODULE_NAM El webservice/index.PHP?v=2&module=ECU STOM_MODULE_NAMEI | GET POST | Custom Module | List of all records |
| webservice/v2/(CUSTOM_MODULE_NAM EMIDATA _ID] webservice/index.PHP?v=2&module=[CU STOM_MODULE_NAME] &data_id=[DATA _ID] | GET PUT | Custom Module | Unique record |
| webservice/v2/(CUSTOM_MODULE_NAM Elicustomfields webservice/index.PHP?v=2&module=[CU STOM_MODULE_NAME184getModuleCust omFields=1 | GET | Custom Module | List of custom fields |

http://[email protected]
AgileBio USA
5473 Kearny Villa Road Suite 255
San Diego, CA 92123
USA
Tel: 347 368 1315
Fax: (800) 453 9128
http://www.agilebio.com
AgileBio Headquarters
75 rue de Lourmel
75015 Paris
FRANCE
Tel: 01 41 79 15 85
Fax: 01 72 70 40 22